- Review
- Open access
- Published:
Long noncoding RNAs in cancer: mechanisms of action and technological advancements
Molecular Cancer volume 15, Article number: 43 (2016)
Abstract
The previous decade has seen long non-coding RNAs (lncRNAs) rise from obscurity to being defined as a category of genetic elements, leaving its mark on the field of cancer biology. With the current number of curated lncRNAs increasing by 10,000 in the last five years, the field is moving from annotation of lncRNA expression in various tumours to understanding their importance in the key cancer signalling networks and characteristic behaviours. Here, we summarize the previously identified as well as recently discovered mechanisms of lncRNA function and their roles in the hallmarks of cancer. Furthermore, we identify novel technologies for investigation of lncRNA properties and their function in carcinogenesis, which will be important for their translation to the clinic as novel biomarkers and therapeutic targets.
Background
Our understanding of cancer biology was drastically changed by the genomic revolution of the last decade, marked by the conclusion of the human genome project and the development of novel DNA sequencing technologies [1, 2]. The complete human genome sequence provided a framework for comparison of populations with cancer susceptibility, allowing for clinical prognosis based on mutations in genes such as BRCA1/2 or differential treatment based modifications in KRAS and BRAF [3–5]. Sequencing of individual tumours revealed the prevalence of acquired DNA damage compared to the germline mutations, which allowed identification of footprints for individual mutagens and gave us important insights into tumour heterogeneity and evolution [6–10]. Parallel to the progress in genomics, advances in transcriptomics initiated functional annotation of numerous genomic loci associated with cancer that do not overlap protein-coding genes – the noncoding genome.
Large-scale cDNA sequencing projects, together with technological advancements such as tiling arrays and the next generation RNA sequencing provided an unprecedented view of the transcriptome complexity [11–15]. Surprisingly, only 1–2 % of the whole genome encodes proteins, with evidence of at least 80 % of the remainder being actively transcribed [11, 16]. These non-coding portions of the genome produce a large variety of mostly regulatory RNAs that differ in their biogenesis, properties and function, and are separated by their size into short, such as miRNAs (reviewed in [17]) and long (>200 nt) RNAs [12, 18–20]. The heterogeneous category of long non-coding RNAs (lncRNA) are especially abundant, accounting for 16,000 curated records in the current Gencode annotation (v.23) [21] with for all lncRNA loci in the human genome numbering as high as 60,000 [22].
lncRNAs remained elusive even in the genomics era due to their low expression levels and their presence in specific cell types, tissues or narrow time frames [23–25]. They were identified as a class of RNA molecules in 2002 [26], even though some lncRNA such as H19 and Xist were known since the early 1990s [27, 28] Analogous to protein coding genes but with low coding potential, these RNAs are usually transcribed by RNA polymerase II (Pol II), spliced, and mostly polyadenylated [12, 13]. Similarly, lncRNA promoters are enriched for active histone modifications typical of Pol II occupancy: H3K4me3, H3K9ac and H3K27ac [20, 29]. Even though the sequence of lncRNAs evolves rapidly, especially compared to their 3D structure, their tissue specificity as well as promotor sequences remain conserved as protein-coding genes [30–32]. The heterogeneity of lncRNAs resonates in the diversity of their functions; lncRNAs interact with DNA, proteins and other RNAs to participate in processes from transcription, intracellular trafficking to chromosome remodelling as reviewed previously [29, 33]).
lncRNAs have been observed to regulate complex cellular behaviours such as growth, differentiation and establishment of cell identity that are commonly deregulated in cancer [34–36]. Some have already been linked to poor prognosis in multiple tumour types and have a clinical relevance as biomarkers. In this review we will focus on the molecular mechanisms of function for cancer-associated lncRNAs, their involvement in cancer hallmarks and provide information on the most recent advances in technologies for their identification and functional interrogation.
Identification of lncRNAs in cancer
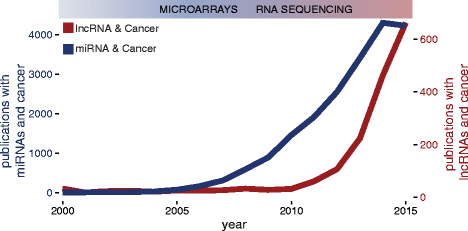
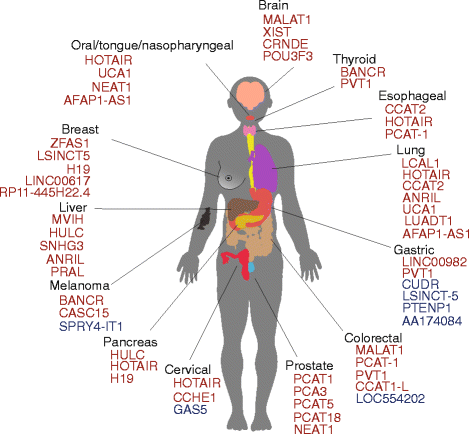
lncRNAs were initially observed in carcinogenesis due to their differential expression compared to normal tissues. High expression in tumour tissues of some of the first identified lncRNAs such as h19, MALAT1 and PCA3 was recognised before the availability of next-generation sequencing technologies [37–39]. RNA sequencing allowed a large-scale assessment of differential expression of lncRNAs comparing cancer to normal tissues, with a large number of lncRNAs showing aberrant expression, similar to the influence microarrays had on the miRNA field (Fig. 1). Recently, a number of lncRNAs have been systematically identified in numerous cancer transcriptomes, either by overlap of sequencing libraries with previously annotated GENCODE lncRNAs, or by de novo assembly of all available public datasets [22, 40], marking their presence in the majority of cancer types (Fig. 2, Additional file 1: Table S1). Novel RNA-seq techniques such as CaptureSeq, that enriches transcript libraries for specific oligonucleotides designed for genomic regions of interest, will improve observation of rare or lowly abundant transcripts from gene deserts associated with cancer [41].
Interest for lncRNAs (red) in the cancer scientific community compared to miRNAs (blue). The y-axis represents the number of publications and the x-axis represents time. Data was obtained by searching Pubmed for ‘lncRNA cancer’ or ‘miRNA cancer’. Data from 2016 was not used in the graph. Publications with terms ‘miRNA’ and ‘cancer’ plateau in 2015
There are two main drivers for altered expression of lncRNAs in tumours. First, abundance of some lncRNAs can be altered due to the cancer-induced change of copy number of their genomic loci. This phenomenon has been observed for FAL1 in ovarian and PRAL in hepatocellular carcinoma [42, 43]. Second, expression of some cancer-associated lncRNAs can be initiated by oncogenes acting as transcription factors. Two crucial genes implicated in multiple tumour types, Myc and p53, act as transcription factors for a large number of lncRNAs [44–46]. Some of these lncRNAs modulate activity of their respective TFs in a feedback loop, for example MINCR with Myc and MEG3 for p53, which makes them potential candidates for therapeutic targeting [47, 48].
Presence of lncRNAs in specific tumours can also be observed based on their overlap with the cancer risk loci identified through genome-wide association studies (GWAS). For example, association of ANRIL with glioma and basal cell carcinoma as well as PTCSC3 with thyroid cancer were discovered based on the known risk loci established through genotyping of cancer patients [49, 50] Recently, CASC15 and NBAT1 were identified through GWAS of neuroblastoma, a cancer that mostly affects children and has a poorly explained genetic background. Both CASC15 and NBAT1 are part of the 6p22 locus that contains SNP rs6939340, associated with metastatic disease and poor event free survival [51, 52]. CASC15 acts as a tumour suppressor and is associated with advanced tumour stages and poor patient survival, while NBAT1 seems to negatively regulate transcription factor NRSF (neuron restrictive silencing factor) [52, 53]. Furthermore, CARLo-5 was identified as significantly correlated with the rs6983267 allele, a single nucleotide polymorphism (SNP) in the region for Myc enhancers. This lncRNA is associated with increased cancer susceptibility and has a function in cell-cycle regulation and tumour development [54]. Since more than 80 % of disease associated SNPs fall into intronic and intergenic regions, it is likely that future development of more sensitive technologies for RNA detection will be essential in defining novel cancer-associated lncRNAs [55].
Mechanisms of lncRNA function in carcinogenesis
Although lncRNAs are scarcely functionally annotated [56], their mechanism of action can be separated based on their influence on chromatin state and methylation, stability of proteins and complexes or by acting as a sponge for miRNA inhibition (reviewed in [29]).
Chromatin state and methylation
Chromatin remodelling was one of the first identified functions of lncRNAs. Epigenetic remodelling is frequently achieved through interaction of a lncRNAs with PRC2 (reviewed in [57]), a protein from the polycomb complex that introduces chromatin inactivation by establishing inhibitory H3K427me3 histone marks [58]. These lncRNAs are sometimes expressed antisense to the target gene involved in cancer, such as in the case of ANRIL with CDKNA/B and asFGFR2 with FGFR2 [59, 60], suggesting a possibility for a cis-acting activity. PCE3, as the first lncRNA involved in cancer was only recently found to have the same mode of action, with the discovery of its antisense protein oncogene PRUNE2 [61]. However, it has also been observed that PRC2 lacks specificity in binding RNAs that recruit it, providing a potential explanation why such a large number of lncRNAs influence chromatin remodelling [62]. Besides acting through PRC2, some lncRNAs such as Kcnq1ot1, TARID, AS1DHRS4 and DACOR1 recruit DNA methyltransferases directly to modify chromatin conformation, or they modify nucleosome positioning through SWI/SNF complex as in the case of SChLAP1 [63–67]. Other lncRNAs, like Firre, bind chromatin remodelers cohesin and CTCF to change chromatin state of the whole chromosomes in the process of X chromosome inactivation [68]. lncRNAs can also act as chromatin activators, as in the case of HOTTIP and CCAT1-L that regulate chromosome looping in their proximity to deposit activating H3K4me3 mark on gene promoters [69–71].
Stability of proteins or protein complexes
A large number of lncRNAs exert their oncogenic function through direct interaction with proteins or protein complexes as scaffolds or allosteric activators/inhibitors. CTBP1-AS, CCTA2 and ZBTB7A interact with transcription factors and modify their activity [72–74]. Some lncRNAs can be used as a scaffold for assembly of whole protein complexes, for example HOTAIR for the HBXIP/Hotair/LSD1 complex [75], NEAT1 for paraspeckle proteins [76, 77], BCAR4 for binding of SNIP1 and PNUTS [78], PRAL for HSP90 and p53 [43]. Furthermore, TERRA recruits a complex of chromatin modifiers to regulate telomere maintenance in response to a variety of cellular signals [79]. Finally, some lncRNAs such as GAS5 bind to nuclear receptors, in this case the glucocorticoid receptor [80].
Competing endogenous RNAs
Some lncRNAs have recently been found to act as competing endogenous RNAs (ceRNAs), by binding miRNAs (“sponging”) and reducing their inhibitory effect on their natural targets (reviewed in [81]). There are numerous examples of lncRNA sponges involved in cancer progression. In lung cancer, UCA1 up-regulates a potent oncogene ERBB4 by binding miR-193-3p [82]. In gastric cancer, MEG3 upregulates Bcl-2 by sequestering miR-181-a [83]. Similarly, ZFAS1 binds miR-150 in hepatocellular carcinoma, Linc-RoR binds miR-145 in endometrial cancer stem cells and CASC2 regulates concentration of miR-21 [84–86].
In addition to the previously described mechanisms, lncRNAs have recently been observed to employ the following set of diverse strategies. lncRNAs can modify the phosphorylation state of proteins by masking phosphorylation motifs, like LINK-a and HIF1a [87]. Furthermore, some lncRNAs such as NORAD act as sponges for a whole set of proteins, in this case the PUMILIO family that would otherwise drive chromosomal instability by repressing mitotic, DNA repair, and DNA replication factors [88]. Interestingly, some lncRNAs form DNA-RNA triplexes that regulate expression of oncogenes either in cis for Khps1, or in trans for MEG3 [89, 90]. Other lncRNAs such as Uc.283 + A control production of miRNAs by influencing processing of the pri-miRNA transcripts, in this case pri-miR-195 [91].
In summary, lncRNAs act through an increasingly wide range of mechanisms that compete with proteins in terms of their diversity and regulatory potential. In the next section we discuss how these mechanisms impact upon cell transformation to cancer phenotype.
Hallmarks of cancer
In 2000, Hanahan and Weinberg described six properties required for cell transformation, termed hallmarks of cancer. These included self-sustained growth signalling, insensitivity to growth inhibition, apoptosis avoidance, uncontrolled proliferation, angiogenesis and metastasis [92, 93]. lncRNAs as regulatory molecules have been implicated in the majority of these functions (reviewed in [94]), and key patterns are starting to emerge.
Self-sustained growth signalling
lncRNAs promote self-sufficiency in growth signals mostly by acting on the activation on the first step of the signal transduction, the signal receptors. Multiple lncRNAs have been observed to specifically bind nuclear receptors either alone or in a ribonucleoprotein complexes (reviewed in [95]). SRA1 serves as a scaffold to stabilize the estrogen receptor [96, 97], while GAS5 acts as the competitive inhibitor of the glucocorticoid receptor [80]. Instead of activating the signal receptors, some lncRNAs such as PVT1 affect the proliferation by regulating the receptor abundance, as is demonstrated for PVT1 and thyroid-stimulating hormone [98].
Insensitivity to growth inhibition
Growth inhibition or its evasion can also be regulated by lncRNAs, and mostly involves influence on tumour suppressors that regulate cell cycle such as cyclins, CDKs, CDK inhibitors and p53 (reviewed in [99]). This can be achieved by chromatin repression through PRC complex, as detailed in the previous section. Using this mechanism PANDA represses protein CDKN1A through PRC1 while ANRIL repress their targets tumour suppressor p15 (CDKN2B) through PRC2 [59, 100, 101]. Some lncRNAs regulate expression of tumour suppressors by influencing various parts of transcription and translation. Initiation of transcription can be influenced by scaffolding of transcription factor complexes, as in the case of LincRNA-p21 and p21 (inhibitor of CDK2) [102]. Transcription elongation can be modified by destabilization of mRNA transcripts, as exemplified by gadd7 and Cdk6 [103]. Finally, transcript stability and translation can be regulated post-transcriptionally by diminishing the role of miRNAs, as in the case of PTENP1 acting as a competitive endogenous RNA to inhibit miRNA repression of PTEN [104]. For some of the lncRNAs like CASC15-S, the direct mechanism of growth inhibition is unknown, but the lack of its expression increases cancer growth and migratory capacity [53].
Avoiding apoptosis
Apoptosis or controlled cell death is one of the key pathways for control of carcinogenesis (reviewed in [36]). Some lncRNAs act on regulation of transcription of key apoptotic genes. For example, lncRNA INXS is expressed from the intron of BCL-X and regulates its splicing into a pro-apoptotic isoform BCL-XS [105]. A recently discovered lncRNA PRAL induces apoptosis by stabilizing a complex between HSP90 and p53 [43]. Several other lncRNAs have been implicated in apoptosis such as SPRY4-IT1 [106], HOXA-AS2 [107] and uc002mbe.2 [108], but the details of the mechanism of action remain still unknown.
Uncontrolled proliferation
Maintenance of telomeres as nucleoprotein structures that stabilize ends of chromosomes is a key requirement for limitless replicative potential of cancer cells. Telomeres shorten in dividing cells, so a ribonucleoprotein complex telomerase is required to elongate telomeric repeats through reverse transcription of an internal template RNA. Shortening of telomeres induces production of a lncRNAs TERRA, that is transcribed from the subtelomeric regions [109]. Under normal conditions, TERRA represses its expression through chromatin modifications, but when activated can recruits protein complexes for homology-directed repair of shortened or damaged telomeric sequences [79].
Promotion of angiogenesis
Multiple lncRNAs have been implicated in regulation of nutrient supply to tumours, mostly by regulating vascular endothelial growth factor (VEGF) that is essential for formation of blood vesicles. Transcription of VEGF was recently reported to be regulated by lncRNAs HOTAIR [110] and MIAT [111]. MIAT sequesters mir-150-5p that is required for repression of VEGF, resulting in microvascular dysfunction and decreased metastasis after MIAT knockdown. MVIH also influences production of VEGF, though indirectly through phosphoglycerate kinase 1 (PGK1) [112]. Finally, MALAT1 has been observed to promotes angiogenic sprouting and migration when expressed in endothelial cells [113].
Tissue invasion and metastasis
Multiple lncRNAs increase invasiveness of cancer cells and facilitate metastasis. Examples of these include h19 [114], MALAT1 in colorectal and nasopharyngeal carcinoma [115], SPRY4-IT1 in melanoma [106], HOTAIR [116], AFAP1-AS1 [117], and CCAT2 [118] in lung cancer, lincRNA-RoR in breast cancer [119], LEIGC in gastric cancer [120] and lncRNA-ATB in hepatocellular carcinoma [121]. Out of these, only lincRNA-RoR and lncRNAs-ATB have a suggested mechanism of action in tissue invasion. lincRNA-RoR likely serves as a “sponge” for miR-145 that is important for regulation of ADP-ribosylation factor 6, a protein involved in invasion of breast cancer cells [119]. Similarly, lncRNA-ATB, acts as a ceRNA to reduce the effect of the miR-200 family targets ZEB1 and ZEB2, two transcription factors that promote cell motility and metastasis [121].
lncRNAs can be involved in a number of other processes related to cancer. Some lncRNAs promote a metabolic switch to glycolysis and lactic acid fermentation termed the Warburg effect [122]. lincRNA-p21 regulates the Warburg effect by preventing ubiquitination of hypoxia-inducible factor-1 (HIF-1), a key transcription factor that promotes upregulation of glycolysis and downregulation of oxidative phosphorylation [123]. Several lncRNAs have been observed as essential for DNA repair by homologous recombination (HR): ANRIL, PCAT1 and DDSR1. Although the mechanism of ANRIL in HR remains unknown, PCAT1 posttranslationally inhibits BRCA2 [124], while DDSR1 is suggested to interact with BRCA1 [125]. Finally, there are implications of lncRNAs on cancer therapies through expression of drug exporters. For example, MRUL promotes expression of ABCB1 that is essential for multidrug-resistance in gastric cancer cell lines [126].
Novel techniques for lncRNA interrogation
The number of annotated transcribed genomic elements has increased by 100 % in the last decade, the majority of which are in the non-coding space and have a defined function in less than 1 % of cases [56]. Such a vast number of novel genetic players presents a great potential for clinical applications, especially in view of cancer as a genomic disease. However, it also requires a thorough rethinking of our basic premises on biological systems, pathway structure and information transfer, as well as a clear technological strategy to identification of their function.
The first challenge is presented by the lack of an exhaustive definition of the full cancer transcriptome, regardless of the cell or tissue type. Currently, a major obstacle to analysis of cancer transcriptome is the alignment of sequence reads to the consensus human genome. Ideally, all the reads would be aligned to a genome sequenced by single-molecule DNA sequencing, but the cost and the quality of this technology are still keeping it away from mainstream research. The next issue is the limited dynamic range of transcript detection for RNA sequencing. This can already be solved by applying the recently developed CaptureSeq method for targeted enrichment of transcripts from specific regions of interest [41]. Furthermore, long read sequencing will be essential for discovery of lncRNAs isoforms and novel exons [127]. In combination with single cell sequencing it will allow identification of individual lncRNAs species from cancer subpopulations, avoiding the heterogeneity of tissue mixture.
After defining the non-coding elements of the transcriptome, the second challenge is the systematic identification of lncRNAs properties that could lead to identifying their cellular function. This can be achieved by investigating their location in the cellular compartments, structural properties as well as possible interactors.
Quantified localization of lncRNAs through microscopy techniques can provide important information about their properties. RNA-Fish as an established technique for RNA localisation has recently been used to identify subcellular location of multiple lncRNAs, in addition to their expression across a population of cells, spatio-temporal behaviour and coexpression with proximal mRNAs [128].
Structure of biological molecules is vital to their function, and several techniques have been developed to investigate secondary and tertiary structures of lncRNAs. Techniques such as Parallel Analysis of RNA Structure (PARS) [129] and Fragmentation Sequencing (FragSeq) [130] sequence RNAs after specific cleavage of single (FragSeq) or single and double stranded (PARS) nucleic acids, allowing for identification of loops in RNA-structure. Another way to investigate structure is to tag the flexible 2′-hydroxyl groups in the RNA backbone by Selective 2′-hydroxyl Acylation and Primer Extension (SHAPE) [131]. Finally, similarly to DNA, RNA can be edited with chemical modifications that modify its structure and binding properties. Two established methods can be used to identify methylated RNA sites: Methylated RNA Immunoprecipitation with next-generation sequencing (MeRIP) [132], or its adaption for hydroxymethylcytosine sites – hMeRIP [133]. Another common RNA-modification is chemical change of nucleotides adenosin to inosin, which can be detected by inosine chemical erasing sequencing (ICE-seq) [134].
Assessing the function of lncRNAs by identifying their binding partners can be performed depending on the type of interaction. Binding of RNA to DNA or proteins can be assessed with ChIRP-seq or ChIRP-MS respectively (Chromatin Isolation by RNA purification followed by sequencing or mass spectrometry) [135, 136]. The specificity of ChIRP is guaranteed by selection of only those RNA that are bound by biotinylated oligonucleotides, similar to RAP [137] and CHART [138], as well as by crosslinking of RNA with DNA or protein by UV or formalin. A recent modification to the protocol can detect individual RNA domains that interact with DNA, RNA or proteins [139]. Instead of biotinylated oligonucleotides, RNA-guided chromatin conformation capture (R3C) reverse-transcribes RNA bound to DNA into cDNA with biotin labelling and joins it with the adjacent genomic DNA with T4 DNA ligase, allowing for streptavidin selection and sequencing [140]. Furthermore, identification of lncRNAs that bind to a protein of interest such as PRC2 [141] can be performed through RNA Immunoprecipitation [142] that was later coupled with sequencing (RIP-Seq) [65]. The specificity of RIP has been improved in by UV crosslinking of RNA and protein in Cross-Linking ImmunoPrecipitation (CLIP) [143] and the later modifications with sequencing (HITS-CLIP) [144] and iClip [145]. Finally, the affinity of a protein for multiple RNA can be assessed in a high-throughput manner. This can be achieved either on a microfluidic platform by RNA-mechanically Induced Trapping of Molecular Interactions (RNA-MITOMI) [146], or on a flow cell in RNA-MaP (massively parallel array) [147].
The final challenge in defining lncRNA functions is developing loss- and gain-of-function lncRNA studies. The RNA interference technology is being supplemented by the powerful CRISPR/Cas-9 system, a newly developed genome-editing technology that allows easier manipulation of lncRNAs behaviour [148]. CRISPR allows multiple types of manipulation, from deletion of various parts of genomic lncRNAs loci, to insertion of promoters, and novel exons. A recent modification of the CRISPR technique that was developed in Rinn group allows insertion of RNA domains to genomic loci, allowing for identification of in cis behaviour of lncRNAs [149].
Conclusion
Long non-coding RNAs are fine-tuners and regulators of key biological processes. Though we have only started to annotate their function in various aspects of cell transformation and metastasis, they are already filling in the major gaps of our understanding of cancer biology. It will be exciting to see the next decade migrate from the perception of lncRNAs as a side act in biological regulation to the center of new biological concepts, paradigms and drug therapies. Watch this space.
Abbreviations
ceRNA, competing endogenous RNAs; CHART, capture hybridization of analysis of RNA targets; ChIRP-seq or ChIRP-MS, chromatin isolation by RNA purification followed by sequencing or mass spectrometry; CLIP-seq, cross-linking immunoprecipitation sequencing; GWAS, genome-wide association studies; hMeRIP, hydroxymethylcytosine sites in RNA; lncRNA, long noncoding RNA; MeRIP, methylation RIP-seq; PARS, parallel analysis of RNA structure; RAP, RNA antisense purification; RIP, RNA immunoprecipitation; RNA-MaP, RNA on a massively parallel array RNA-MITOMI, RNA-mechanically induced trapping of molecular interactions; SHAPE, selective 2′-hydroxyl acylation and primer extension; SNP, single nucleotide polymorphism
References
Wheeler DA, Wang L. From human genome to cancer genome: the first decade. Genome Res. 2013;23:1054–62.
Offit K. Decade in review--genomics: a decade of discovery in cancer genomics. Nat Rev Clin Oncol. 2014;11:632–4.
Pal T, Permuth-Wey J, Betts JA, Krischer JP, Fiorica J, Arango H, LaPolla J, Hoffman M, Martino MA, Wakeley K, Wilbanks G, Nicosia S, Cantor A, Sutphen R. BRCA1 and BRCA2 mutations account for a large proportion of ovarian carcinoma cases. Cancer. 2005;104:2807–16.
Lievre A, Bachet J-B, Le Corre D, Boige V, Landi B, Emile J-F, Cote J-F, Tomasic G, Penna C, Ducreux M, Rougier P, Penault-Llorca F, Laurent-Puig P. KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res. 2006;66:3992–5.
Zlobec I, Bihl MP, Schwarb H, Terracciano L, Lugli A. Clinicopathological and protein characterization of BRAF- and K-RAS-mutated colorectal cancer and implications for prognosis. Int J Cancer. 2010;127:367–80.
Forbes SA, Beare D, Gunasekaran P, Leung K, Bindal N, Boutselakis H, Ding M, Bamford S, Cole C, Ward S, Kok CY, Jia M, De T, Teague JW, Stratton MR, McDermott U, Campbell PJ. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 2015;43(Database issue):D805–11.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SAJR, Behjati S, Biankin AV, Bignell GR, Bolli N, Borg A, Børresen-Dale A-L, Boyault S, Burkhardt B, Butler AP, Caldas C, Davies HR, Desmedt C, Eils R, Eyfjörd JE, Foekens JA, Greaves M, Hosoda F, Hutter B, Ilicic T, Imbeaud S, Imielinski M, Jager N, Jones DTW, Jones D, Knappskog S, Kool M, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Campbell PJ, Pleasance ED, Stephens PJ, Dicks E, Rance R, Goodhead I, Follows GA, Green AR, Futreal PA, Stratton MR. Subclonal phylogenetic structures in cancer revealed by ultra-deep sequencing. Proc Natl Acad Sci U S A. 2008;105:13081–6.
Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A, Carter SL, Stewart C, Mermel CH, Roberts SA, Kiezun A, Hammerman PS, McKenna A, Drier Y, Zou L, Ramos AH, Pugh TJ, Stransky N, Helman E, Kim J, Sougnez C, Ambrogio L, Nickerson E, Shefler E, Cortés ML, Auclair D, Saksena G, Voet D, Noble M, DiCara D, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013;499:214–8.
Navin N, Kendall J, Troge J, Andrews P, Rodgers L, McIndoo J, Cook K, Stepansky A, Levy D, Esposito D, Muthuswamy L, Krasnitz A, McCombie WR, Hicks J, Wigler M. Tumour evolution inferred by single-cell sequencing. Nature. 2011;472:90–4.
Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74.
Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, Brenner SE, Batalov S, Forrest ARR, Zavolan M, Davis MJ, Wilming LG, Aidinis V, Allen JE, Ambesi-Impiombato A, Apweiler R, Aturaliya RN, Bailey TL, Bansal M, Baxter L, Beisel KW, Bersano T, Bono H, et al. The transcriptional landscape of the mammalian genome. Science. 2005;309:1559–63.
Bertone P, Stolc V, Royce TE, Rozowsky JS, Urban AE, Zhu X, Rinn JL, Tongprasit W, Samanta M, Weissman S, Gerstein M, Snyder M. Global identification of human transcribed sequences with genome tiling arrays. Science. 2004;306:2242–6.
Dinger ME, Amaral PP, Mercer TR, Mattick JS. Pervasive transcription of the eukaryotic genome: functional indices and conceptual implications. Brief Funct Genomic Proteomic. 2009;8:407–23.
Clark MB, Amaral PP, Schlesinger FJ, Dinger ME, Taft RJ, Rinn JL, Ponting CP, Stadler PF, Morris KV, Morillon A, Rozowsky JS, Gerstein MB, Wahlestedt C, Hayashizaki Y, Carninci P, Gingeras TR, Mattick JS. The reality of pervasive transcription. PLoS Biol. 2011;9:e1000625. discussion e1001102.
Hangauer MJ, Vaughn IW, McManus MT. Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS Genet. 2013;9:e1003569.
Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer. 2015;15:321–33.
Engstrom PG, Suzuki H, Ninomiya N, Akalin A, Sessa L, Lavorgna G, Brozzi A, Luzi L, Tan SL, Yang L, Kunarso G, Ng EL-C, Batalov S, Wahlestedt C, Kai C, Kawai J, Carninci P, Hayashizaki Y, Wells C, Bajic VB, Orlando V, Reid JF, Lenhard B, Lipovich L. Complex Loci in human and mouse genomes. PLoS Genet. 2006;2:e47.
Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermuller J, Hofacker IL, Bell I, Cheung E, Drenkow J, Dumais E, Patel S, Helt G, Ganesh M, Ghosh S, Piccolboni A, Sementchenko V, Tammana H, Gingeras TR. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007;316:1484–8.
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, Jacks T, Hacohen N, Bernstein BE, Kellis M, Regev A, Rinn JL, Lander ES. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458:223–7.
Harrow J, Frankish A, Gonzalez JM, Tapanari E, Diekhans M, Kokocinski F, Aken BL, Barrell D, Zadissa A, Searle S, Barnes I, Bignell A, Boychenko V, Hunt T, Kay M, Mukherjee G, Rajan J, Despacio-Reyes G, Saunders G, Steward C, Harte R, Lin M, Howald C, Tanzer A, Derrien T, Chrast J, Walters N, Balasubramanian S, Pei B, Tress M, et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 2012;22:1760–74.
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, Barrette TR, Prensner JR, Evans JR, Zhao S, Poliakov A, Cao X, Dhanasekaran SM, Wu Y-M, Robinson DR, Beer DG, Feng FY, Iyer HK, Chinnaiyan AM. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47:199–208.
Mercer TR, Dinger ME, Sunkin SM, Mehler MF, Mattick JS. Specific expression of long noncoding RNAs in the mouse brain. Proc Natl Acad Sci U S A. 2008;105:716–21.
Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011;25:1915–27.
Gloss BS, Dinger ME. The specificity of long noncoding RNA expression. Biochim Biophys Acta. 1859;2016:16–22.
Okazaki Y, Furuno M, Kasukawa T, Adachi J, Bono H, Kondo S, Nikaido I, Osato N, Saito R, Suzuki H, Yamanaka I, Kiyosawa H, Yagi K, Tomaru Y, Hasegawa Y, Nogami A, Schonbach C, Gojobori T, Baldarelli R, Hill DP, Bult C, Hume DA, Quackenbush J, Schriml LM, Kanapin A, Matsuda H, Batalov S, Beisel KW, Blake JA, Bradt D, et al. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nat Genet. 2002;420:563–73.
Brannan CI, Dees EC, Ingram RS, Tilghman SM. The product of the H19 gene may function as an RNA. Mol Cell Biol. 1990;10:28–36.
Brockdorff N, Ashworth A, Kay GF, McCabe VM, Norris DP, Cooper PJ, Swift S, Rastan S. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell. 1992;71:515–26.
Quinn JJ, Chang HY. Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet. 2015;17:47–62.
Smith MA, Gesell T, Stadler PF, Mattick JS. Widespread purifying selection on RNA structure in mammals. Nucleic Acids Res. 2013;41:8220–36.
Rands CM, Meader S, Ponting CP, Lunter G. 8.2 % of the human genome is constrained: variation in rates of turnover across functional element classes in the human lineage. PLoS Genet. 2014;10:e1004525.
Necsulea A, Soumillon M, Warnefors M, Liechti A, Daish T, Zeller U, Baker JC, Grutzner F, Kaessmann H. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature. 2014;505:635.
Rinn JL, Chang HY. Genome regulation by long noncoding RNAs. Annu Rev Biochem. 2012;81:145–66.
Hu W, Alvarez-Dominguez JR, Lodish HF. Regulation of mammalian cell differentiation by long non-coding RNAs. EMBO Rep. 2012;13:971–83.
Flynn RA, Chang HY. Long noncoding RNAs in cell-fate programming and reprogramming. Cell Stem Cell. 2014;14:752–61.
Rossi MN, Antonangeli F. LncRNAs: new players in apoptosis control. Int J Cell Biol. 2014;2014:473857.
Zhang Y, Shields T, Crenshaw T, Hao Y, Moulton T, Tycko B. Imprinting of human H19: allele-specific CpG methylation, loss of the active allele in Wilms tumor, and potential for somatic allele switching. Am J Hum Genet. 1993;53:113–24.
Luo J-H, Ren B, Keryanov S, Tseng GC, Rao UNM, Monga SP, Strom S, Demetris AJ, Nalesnik M, Yu YP, Ranganathan S, Michalopoulos GK. Transcriptomic and genomic analysis of human hepatocellular carcinomas and hepatoblastomas. Hepatology. 2006;44:1012–24.
de Kok JB, Verhaegh GW, Roelofs RW, Hessels D, Kiemeney LA, Aalders TW, Swinkels DW, Schalken JA. DD3(PCA3), a very sensitive and specific marker to detect prostate tumors. Cancer Res. 2002;62:2695–8.
Yan X, Hu Z, Feng Y, Hu X, Yuan J, Zhao SD, Zhang Y, Yang L, Shan W, He Q, Fan L, Kandalaft LE, Tanyi JL, Li C, Yuan C-X, Zhang D, Yuan H, Hua K, Lu Y, Katsaros D, Huang Q, Montone K, Fan Y, Coukos G, Boyd J, Sood AK, Rebbeck T, Mills GB, Dang CV, Zhang L. Comprehensive genomic characterization of long non-coding RNAs across human cancers. Cancer Cell. 2015;28:529–40.
Clark MB, Mercer TR, Bussotti G, Leonardi T, Haynes KR, Crawford J, Brunck ME, Cao K-AL, Thomas GP, Chen WY, Taft RJ, Nielsen LK, Enright AJ, Mattick JS, Dinger ME. Quantitative gene profiling of long noncoding RNAs with targeted RNA sequencing. Nat Meth. 2015;12:339–42.
Hu X, Feng Y, Zhang D, Zhao SD, Hu Z, Greshock J, Zhang Y, Yang L, Zhong X, Wang L-P, Jean S, Li C, Huang Q, Katsaros D, Montone KT, Tanyi JL, Lu Y, Boyd J, Nathanson KL, Li H, Mills GB, Zhang L. A functional genomic approach identifies FAL1 as an oncogenic long noncoding RNA that associates with BMI1 and represses p21 expression in cancer. Cancer Cell. 2014;26:344–57.
Zhou C-C, Yang F, Yuan S-X, Ma J-Z, Liu F, Yuan J-H, Bi F-R, Lin K-Y, Yin J-H, Cao G-W, Zhou W-P, Wang F, Sun S-H. Systemic genome screening identified the outcome associated focal loss of long noncoding RNA PRAL in hepatocellular carcinoma. Hepatology. 2015;63:850.
Hart JR, Roberts TC, Weinberg MS, Morris KV, Vogt PK. MYC regulates the non-coding transcriptome. Oncotarget. 2014;5:12543–54.
Younger ST, Kenzelmann-Broz D, Jung H, Attardi LD, Rinn JL. Integrative genomic analysis reveals widespread enhancer regulation by p53 in response to DNA damage. Nucleic Acids Res. 2015;43:4447–62.
Grossi E, Sanchez Y, Huarte M. Expanding the p53 regulatory network: LncRNAs take up the challenge. Biochim Biophys Acta. 1859;2016:200–8.
Doose G, Haake A, Bernhart SH, Lopez C, Duggimpudi S, Wojciech F, Bergmann AK, Borkhardt A, Burkhardt B, Claviez A, Dimitrova L, Haas S, Hoell JI, Hummel M, Karsch D, Klapper W, Kleo K, Kretzmer H, Kreuz M, Kuppers R, Lawerenz C, Lenze D, Loeffler M, Mantovani-Loffler L, Moller P, Ott G, Richter J, Rohde M, Rosenstiel P, Rosenwald A, et al. MINCR is a MYC-induced lncRNA able to modulate MYC’s transcriptional network in Burkitt lymphoma cells. Proc Natl Acad Sci U S A. 2015;112:E5261–70.
Li J, Bian E-B, He X-J, Ma C-C, Zong G, Wang H-L, Zhao B. Epigenetic repression of long non-coding RNA MEG3 mediated by DNMT1 represses the p53 pathway in gliomas. Int J Oncol. 2016;48:723–33.
Pasmant E, Sabbagh A, Masliah-Planchon J, Ortonne N, Laurendeau I, Melin L, Ferkal S, Hernandez L, Leroy K, Valeyrie-Allanore L, Parfait B, Vidaud D, Bieche I, Lantieri L, Wolkenstein P, Vidaud M. Role of noncoding RNA ANRIL in genesis of plexiform neurofibromas in neurofibromatosis type 1. J Natl Cancer Inst. 2011;103:1713–22.
Jendrzejewski J, He H, Radomska HS, Li W, Tomsic J, Liyanarachchi S, Davuluri RV, Nagy R, la Chapelle de A. The polymorphism rs944289 predisposes to papillary thyroid carcinoma through a large intergenic noncoding RNA gene of tumor suppressor type. Proc Natl Acad Sci U S A. 2012;109:8646–51.
Diskin SJ, Capasso M, Schnepp RW, Cole KA, Attiyeh EF, Hou C, Diamond M, Carpenter EL, Winter C, Lee H, Jagannathan J, Latorre V, Iolascon A, Hakonarson H, Devoto M, Maris JM. Common variation at 6q16 within HACE1 and LIN28B influences susceptibility to neuroblastoma. Nat Genet. 2012;44:1126–30.
Pandey GK, Mitra S, Subhash S, Hertwig F, Kanduri M, Mishra K, Fransson S, Ganeshram A, Mondal T, Bandaru S, Ostensson M, Akyurek LM, Abrahamsson J, Pfeifer S, Larsson E, Shi L, Peng Z, Fischer M, Martinsson T, Hedborg F, Kogner P, Kanduri C. The risk-associated long noncoding RNA NBAT-1 controls neuroblastoma progression by regulating cell proliferation and neuronal differentiation. Cancer Cell. 2014;26:722–37.
Russell MR, Penikis A, Oldridge DA, Alvarez-Dominguez JR, McDaniel L, Diamond M, Padovan O, Raman P, Li Y, Wei JS, Zhang S, Gnanchandran J, Seeger R, Asgharzadeh S, Khan J, Diskin SJ, Maris JM, Cole KA. CASC15-S Is a Tumor Suppressor lncRNA at the 6p22 Neuroblastoma Susceptibility Locus. Cancer Res. 2015;75:3155–66.
Kim T, Cui R, Jeon Y-J, Lee J-H, Lee JH, Sim H, Park JK, Fadda P, Tili E, Nakanishi H, Huh M-I, Kim S-H, Cho JH, Sung BH, Peng Y, Lee TJ, Luo Z, Sun H-L, Wei H, Alder H, Oh JS, Shim KS, Ko S-B, Croce CM. Long-range interaction and correlation between MYC enhancer and oncogenic long noncoding RNA CARLo-5. Proc Natl Acad Sci U S A. 2014;111:4173–8.
Welter D, MacArthur J, Morales J, Burdett T, Hall P, Junkins H, Klemm A, Flicek P, Manolio T, Hindorff L, Parkinson H. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 2014;42(Database issue):D1001–6.
Quek XC, Thomson DW, Maag JLV, Bartonicek N, Signal B, Clark MB, Gloss BS, Dinger ME. lncRNAdb v2.0: expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res. 2015;43(Database issue):D168–73.
Davidovich C, Cech TR. The recruitment of chromatin modifiers by long noncoding RNAs: lessons from PRC2. RNA. 2015;21:2007–22.
Ringrose L, Ehret H, Paro R. Distinct contributions of histone H3 lysine 9 and 27 methylation to locus-specific stability of polycomb complexes. Mol Cell. 2004;16:641–53.
Kotake Y, Nakagawa T, Kitagawa K, Suzuki S, Liu N, Kitagawa M, Xiong Y. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene. 2011;30:1956–62.
Gonzalez I, Munita R, Agirre E, Dittmer TA, Gysling K, Misteli T, Luco RF. A lncRNA regulates alternative splicing via establishment of a splicing-specific chromatin signature. Nat Struct Mol Biol. 2015;22:370–6.
Salameh A, Lee AK, Cardo-Vila M, Nunes DN, Efstathiou E, Staquicini FI, Dobroff AS, Marchio S, Navone NM, Hosoya H, Lauer RC, Wen S, Salmeron CC, Hoang A, Newsham I, Lima LA, Carraro DM, Oliviero S, Kolonin MG, Sidman RL, Do K-A, Troncoso P, Logothetis CJ, Brentani RR, Calin GA, Cavenee WK, Dias-Neto E, Pasqualini R, Arap W. PRUNE2 is a human prostate cancer suppressor regulated by the intronic long noncoding RNA PCA3. Proc Natl Acad Sci U S A. 2015;112:8403–8.
Davidovich C, Wang X, Cifuentes-Rojas C, Goodrich KJ, Gooding AR, Lee JT, Cech TR. Toward a consensus on the binding specificity and promiscuity of PRC2 for RNA. Mol Cell. 2015;57:552–8.
Mohammad F, Mondal T, Guseva N, Pandey GK, Kanduri C. Kcnq1ot1 noncoding RNA mediates transcriptional gene silencing by interacting with Dnmt1. Development. 2010;137:2493–9.
Li Q, Su Z, Xu X, Liu G, Song X, Wang R, Sui X, Liu T, Chang X, Huang D. AS1DHRS4, a head-to-head natural antisense transcript, silences the DHRS4 gene cluster in cis and trans. Proc Natl Acad Sci U S A. 2012;109:14110–5.
Merry CR, Forrest ME, Sabers JN, Beard L, Gao X-H, Hatzoglou M, Jackson MW, Wang Z, Markowitz SD, Khalil AM. DNMT1-associated long non-coding RNAs regulate global gene expression and DNA methylation in colon cancer. Hum Mol Genet. 2015;24:6240–53.
Lee RS, Roberts CWM. Linking the SWI/SNF complex to prostate cancer. Nat Genet. 2013;45:1268–9.
Arab K, Park YJ, Lindroth AM, Schafer A, Oakes C, Weichenhan D, Lukanova A, Lundin E, Risch A, Meister M, Dienemann H, Dyckhoff G, Herold-Mende C, Grummt I, Niehrs C, Plass C. Long noncoding RNA TARID directs demethylation and activation of the tumor suppressor TCF21 via GADD45A. Mol Cell. 2014;55:604–14.
Yang F, Deng X, Ma W, Berletch JB, Rabaia N, Wei G, Moore JM, Filippova GN, Xu J, Liu Y, Noble WS, Shendure J, Disteche CM. The lncRNA Firre anchors the inactive X chromosome to the nucleolus by binding CTCF and maintains H3K27me3 methylation. Genome Biol. 2015;16:52.
Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA, Gupta RA, Wysocka J, Lei M, Dekker J, Helms JA, Chang HY. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature. 2011;472:120–4.
Lai F, Orom UA, Cesaroni M, Beringer M, Taatjes DJ, Blobel GA, Shiekhattar R. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature. 2013;494:497–501.
Xiang J-F, Yin Q-F, Chen T, Zhang Y, Zhang X-O, Wu Z, Zhang S, Wang H-B, Ge J, Lu X, Yang L, Chen L-L. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 2014;24:513–31.
Takayama K-I, Horie-Inoue K, Katayama S, Suzuki T, Tsutsumi S, Ikeda K, Urano T, Fujimura T, Takagi K, Takahashi S, Homma Y, Ouchi Y, Aburatani H, Hayashizaki Y, Inoue S. Androgen-responsive long noncoding RNA CTBP1-AS promotes prostate cancer. EMBO J. 2013;32:1665–80.
Ling H, Spizzo R, Atlasi Y, Nicoloso M, Shimizu M, Redis RS, Nishida N, Gafa R, Song J, Guo Z, Ivan C, Barbarotto E, De Vries I, Zhang X, Ferracin M, Churchman M, van Galen JF, Beverloo BH, Shariati M, Haderk F, Estecio MR, Garcia-Manero G, Patijn GA, Gotley DC, Bhardwaj V, Shureiqi I, Sen S, Multani AS, Welsh J, Yamamoto K, et al. CCAT2, a novel noncoding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res. 2013;23:1446–61.
Wang G, Lunardi A, Zhang J, Chen Z, Ala U, Webster KA, Tay Y, Gonzalez-Billalabeitia E, Egia A, Shaffer DR, Carver B, Liu X-S, Taulli R, Kuo WP, Nardella C, Signoretti S, Cordon-Cardo C, Gerald WL, Pandolfi PP. Zbtb7a suppresses prostate cancer through repression of a Sox9-dependent pathway for cellular senescence bypass and tumor invasion. Nat Genet. 2013;45:739–46.
Li Y, Wang Z, Shi H, Li H, Li L, Fang R, Cai X, Liu B, Zhang X, Ye L. HBXIP and LSD1 Scaffolded by lncRNA hotair mediate transcriptional activation by c-Myc. Cancer Res. 2015;76:293.
Clemson CM, Hutchinson JN, Sara SA, Ensminger AW, Fox AH, Chess A, Lawrence JB. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol Cell. 2009;33:717–26.
Naganuma T, Nakagawa S, Tanigawa A, Sasaki YF, Goshima N, Hirose T. Alternative 3′-end processing of long noncoding RNA initiates construction of nuclear paraspeckles. EMBO J. 2012;31:4020–34.
Xing Z, Lin A, Li C, Liang K, Wang S, Liu Y, Park PK, Qin L, Wei Y, Hawke DH, Hung M-C, Lin C, Yang L. lncRNA directs cooperative epigenetic regulation downstream of chemokine signals. Cell. 2014;159:1110–25.
Rippe K, Luke B. TERRA and the state of the telomere. Nat Struct Mol Biol. 2015;22:853–8.
Mourtada-Maarabouni M, Pickard MR, Hedge VL, Farzaneh F, Williams GT. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene. 2009;28:195–208.
Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014;505:344–52.
Nie W, Ge H-J, Yang X-Q, Sun X, Huang H, Tao X, Chen W-S, Li B. LncRNA-UCA1 exerts oncogenic functions in non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett. 2016;371:99–106.
Peng W, Si S, Zhang Q, Li C, Zhao F, Wang F, Yu J, Ma R. Long non-coding RNA MEG3 functions as a competing endogenous RNA to regulate gastric cancer progression. J Exp Clin Cancer Res. 2015;34:79.
Deng L, Yang S-B, Xu F-F, Zhang J-H. Long noncoding RNA CCAT1 promotes hepatocellular carcinoma progression by functioning as let-7 sponge. J Exp Clin Cancer Res. 2015;34:18.
Zhou X, Gao Q, Wang J, Zhang X, Liu K, Duan Z. Linc-RNA-RoR acts as a “sponge” against mediation of the differentiation of endometrial cancer stem cells by microRNA-145. Gynecol Oncol. 2014;133:333–9.
Wang P, Liu Y-H, Yao Y-L, Li Z, Li Z-Q, Ma J, Xue Y-X. Long non-coding RNA CASC2 suppresses malignancy in human gliomas by miR-21. Cell Signal. 2015;27:275–82.
Liu B, Sun L, Liu Q, Gong C, Yao Y, Lv X, Lin L, Yao H, Su F, Li D, Zeng M, Song E. A cytoplasmic NF-kappaB interacting long noncoding RNA blocks IkappaB phosphorylation and suppresses breast cancer metastasis. Cancer Cell. 2015;27:370–81.
Lee S, Kopp F, Chang T-C, Sataluri A, Chen B, Sivakumar S, Yu H, Xie Y, Mendell JT. Noncoding RNA NORAD regulates genomic stability by sequestering PUMILIO proteins. Cell. 2015;164:69.
Postepska-Igielska A, Giwojna A, Gasri-Plotnitsky L, Schmitt N, Dold A, Ginsberg D, Grummt I. LncRNA Khps1 regulates expression of the proto-oncogene SPHK1 via triplex-mediated changes in chromatin structure. Mol Cell. 2015;60:626–36.
Mondal T, Subhash S, Vaid R, Enroth S, Uday S, Reinius B, Mitra S, Mohammed A, James AR, Hoberg E, Moustakas A, Gyllensten U, Jones SJM, Gustafsson CM, Sims AH, Westerlund F, Gorab E, Kanduri C. MEG3 long noncoding RNA regulates the TGF-beta pathway genes through formation of RNA-DNA triplex structures. Nat Commun. 2015;6:7743.
Liz J, Portela A, Soler M, Gomez A, Ling H, Michlewski G, Calin GA, Guil S, Esteller M. Regulation of pri-miRNA processing by a long noncoding RNA transcribed from an ultraconserved region. Mol Cell. 2014;55:138–47.
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57–70.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Gutschner T, Diederichs S. The hallmarks of cancer: a long non-coding RNA point of view. RNA Biol. 2012;9:703–19.
Cathcart P, Lucchesi W, Ottaviani S, De Giorgio A, Krell J, Stebbing J, Castellano L. Noncoding RNAs and the control of signalling via nuclear receptor regulation in health and disease. Best Pract Res Clin Endocrinol Metab. 2015;29:529–43.
Lanz RB, McKenna NJ, Onate SA, Albrecht U, Wong J, Tsai SY, Tsai MJ, O’Malley BW. A steroid receptor coactivator, SRA, functions as an RNA and is present in an SRC-1 complex. Cell. 1999;97:17–27.
Shi Y, Downes M, Xie W, Kao HY, Ordentlich P, Tsai CC, Hon M, Evans RM. Sharp, an inducible cofactor that integrates nuclear receptor repression and activation. Genes Dev. 2001;15:1140–51.
Zhou Q, Chen J, Feng J, Wang J. Long noncoding RNA PVT1 modulates thyroid cancer cell proliferation by recruiting EZH2 and regulating thyroid-stimulating hormone receptor (TSHR). Tumour Biol. 2015.
Kitagawa M, Kitagawa K, Kotake Y, Niida H, Ohhata T. Cell cycle regulation by long non-coding RNAs. Cell Mol Life Sci. 2013;70:4785–94.
Puvvula PK, Desetty RD, Pineau P, Marchio A, Moon A, Dejean A, Bischof O. Long noncoding RNA PANDA and scaffold-attachment-factor SAFA control senescence entry and exit. Nat Commun. 2014;5:5323.
Benatti P, Belluti S, Miotto B, Neusiedler J, Dolfini D, Drac M, Basile V, Schwob E, Mantovani R, Julian Blow J, Imbriano C. Direct non transcriptional role of NF-Y in DNA replication. Biochim Biophys Acta. 2015;1863:673.
Dimitrova N, Zamudio JR, Jong RM, Soukup D, Resnick R, Sarma K, Ward AJ, Raj A, Lee JT, Sharp PA, Jacks T. LincRNA-p21 activates p21 in cis to promote Polycomb target gene expression and to enforce the G1/S checkpoint. Mol Cell. 2014;54:777–90.
Liu X, Li D, Zhang W, Guo M, Zhan Q. Long non-coding RNA gadd7 interacts with TDP-43 and regulates Cdk6 mRNA decay. EMBO J. 2012;31:4415–27.
Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature. 2010;465:1033–8.
DeOcesano-Pereira C, Amaral MS, Parreira KS, Ayupe AC, Jacysyn JF, Amarante-Mendes GP, Reis EM, Verjovski-Almeida S. Long non-coding RNA INXS is a critical mediator of BCL-XS induced apoptosis. Nucleic Acids Res. 2014;42:8343–55.
Mazar J, Zhao W, Khalil AM, Lee B, Shelley J, Govindarajan SS, Yamamoto F, Ratnam M, Aftab MN, Collins S, Finck BN, Han X, Mattick JS, Dinger ME, Perera RJ. The functional characterization of long noncoding RNA SPRY4-IT1 in human melanoma cells. Oncotarget. 2014;5:8959–69.
Zhao H, Zhang X, Frazao JB, Condino-Neto A, Newburger PE. HOX antisense lincRNA HOXA-AS2 is an apoptosis repressor in all trans retinoic acid treated NB4 promyelocytic leukemia cells. J Cell Biochem. 2013;114:2375–83.
Yang H, Zhong Y, Xie H, Lai X, Xu M, Nie Y, Liu S, Wan Y-JY. Induction of the liver cancer-down-regulated long noncoding RNA uc002mbe.2 mediates trichostatin-induced apoptosis of liver cancer cells. Biochem Pharmacol. 2013;85:1761–9.
Redon S, Reichenbach P, Lingner J. The non-coding RNA TERRA is a natural ligand and direct inhibitor of human telomerase. Nucleic Acids Res. 2010;38:5797–806.
Fu W-M, Lu Y-F, Hu B-G, Liang W-C, Zhu X, Yang H-D, Li G, Zhang J-F. Long noncoding RNA hotair mediated angiogenesis in nasopharyngeal carcinoma by direct and indirect signaling pathways. Oncotarget. 2015;7:4712.
Yan B, Yao J, Liu J-Y, Li X-M, Wang X-Q, Li Y-J, Tao Z-F, Song Y-C, Chen Q, Jiang Q. lncRNA-MIAT regulates microvascular dysfunction by functioning as a competing endogenous RNA. Circ Res. 2015;116:1143–56.
Yuan S-X, Yang F, Yang Y, Tao Q-F, Zhang J, Huang G, Yang Y, Wang R-Y, Yang S, Huo X-S, Zhang L, Wang F, Sun S-H, Zhou W-P. Long noncoding RNA associated with microvascular invasion in hepatocellular carcinoma promotes angiogenesis and serves as a predictor for hepatocellular carcinoma patients’ poor recurrence-free survival after hepatectomy. Hepatology. 2012;56:2231–41.
Michalik KM, You X, Manavski Y, Doddaballapur A, Zornig M, Braun T, John D, Ponomareva Y, Chen W, Uchida S, Boon RA, Dimmeler S. Long noncoding RNA MALAT1 regulates endothelial cell function and vessel growth. Circ Res. 2014;114:1389–97.
Raveh E, Matouk IJ, Gilon M, Hochberg A. The H19 Long non-coding RNA in cancer initiation, progression and metastasis - a proposed unifying theory. Mol Cancer. 2015;14:184.
Yang M-H, Hu Z-Y, Xu C, Xie L-Y, Wang X-Y, Chen S-Y, Li Z-G. MALAT1 promotes colorectal cancer cell proliferation/migration/invasion via PRKA kinase anchor protein 9. Biochim Biophys Acta. 2015;1852:166–74.
Arun G, Diermeier S, Akerman M, Chang K-C, Wilkinson JE, Hearn S, Kim Y, MacLeod AR, Krainer AR, Norton L, Brogi E, Egeblad M, Spector DL. Differentiation of mammary tumors and reduction in metastasis upon Malat1 lncRNA loss. Genes Dev. 2016;30:34–51.
Deng J, Liang Y, Liu C, He S, Wang S. The up-regulation of long non-coding RNA AFAP1-AS1 is associated with the poor prognosis of NSCLC patients. Biomed Pharmacother. 2015;75:8–11.
Qiu M, Xu Y, Yang X, Wang J, Hu J, Xu L, Yin R. CCAT2 is a lung adenocarcinoma-specific long non-coding RNA and promotes invasion of non-small cell lung cancer. Tumour Biol. 2014;35:5375–80.
Eades G, Wolfson B, Zhang Y, Li Q, Yao Y, Zhou Q. lincRNA-RoR and miR-145 regulate invasion in triple-negative breast cancer via targeting ARF6. Mol Cancer Res. 2015;13:330–8.
Han Y, Ye J, Wu D, Wu P, Chen Z, Chen J, Gao S, Huang J. LEIGC long non-coding RNA acts as a tumor suppressor in gastric carcinoma by inhibiting the epithelial-to-mesenchymal transition. BMC Cancer. 2014;14:932.
Yuan J-H, Yang F, Wang F, Ma J-Z, Guo Y-J, Tao Q-F, Liu F, Pan W, Wang T-T, Zhou C-C, Wang S-B, Wang Y-Z, Yang Y, Yang N, Zhou W-P, Yang G-S, Sun S-H. A long noncoding RNA activated by TGF-beta promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell. 2014;25:666–81.
Gatenby RA, Gillies RJ. Why do cancers have high aerobic glycolysis? Nat Rev Cancer. 2004;4:891–9.
Yang F, Zhang H, Mei Y, Wu M. Reciprocal regulation of HIF-1alpha and lincRNA-p21 modulates the Warburg effect. Mol Cell. 2014;53:88–100.
Prensner JR, Chen W, Iyer MK, Cao Q, Ma T, Han S, Sahu A, Malik R, Wilder-Romans K, Navone N, Logothetis CJ, Araujo JC, Pisters LL, Tewari AK, Canman CE, Knudsen KE, Kitabayashi N, Rubin MA, Demichelis F, Lawrence TS, Chinnaiyan AM, Feng FY. PCAT-1, a long noncoding RNA, regulates BRCA2 and controls homologous recombination in cancer. Cancer Res. 2014;74:1651–60.
Sharma V, Khurana S, Kubben N, Abdelmohsen K, Oberdoerffer P, Gorospe M, Misteli T. A BRCA1-interacting lncRNA regulates homologous recombination. EMBO Rep. 2015;16:1520–34.
Wang Y, Zhang D, Wu K, Zhao Q, Nie Y, Fan D. Long noncoding RNA MRUL promotes ABCB1 expression in multidrug-resistant gastric cancer cell sublines. Mol Cell Biol. 2014;34:3182–93.
Rhoads A, Au KF. PacBio sequencing and its applications. Genomics Proteomics Bioinformatics. 2015;13:278–89.
Cabili MN, Dunagin MC, McClanahan PD, Biaesch A, Padovan-Merhar O, Regev A, Rinn JL, Raj A. Localization and abundance analysis of human lncRNAs at single-cell and single-molecule resolution. Genome Biol. 2015;16:20.
Kertesz M, Wan Y, Mazor E, Rinn JL, Nutter RC, Chang HY, Segal E. Genome-wide measurement of RNA secondary structure in yeast. Nature. 2010;467:103–7.
Underwood JG, Uzilov AV, Katzman S, Onodera CS, Mainzer JE, Mathews DH, Lowe TM, Salama SR, Haussler D. FragSeq: transcriptome-wide RNA structure probing using high-throughput sequencing. Nat Meth. 2010;7:995–1001.
Merino EJ, Wilkinson KA, Coughlan JL, Weeks KM. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J Am Chem Soc. 2005;127:4223–31.
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell. 2012;149:1635–46.
Delatte B, Wang F, Ngoc LV, Collignon E, Bonvin E, Deplus R, Calonne E, Hassabi B, Putmans P, Awe S, Wetzel C, Kreher J, Soin R, Creppe C, Limbach PA, Gueydan C, Kruys V, Brehm A, Minakhina S, Defrance M, Steward R, Fuks F. Transcriptome-wide distribution and function of RNA hydroxymethylcytosine. Science. 2016;351:282–5.
Suzuki T, Ueda H, Okada S, Sakurai M. Transcriptome-wide identification of adenosine-to-inosine editing using the ICE-seq method. Nat Protoc. 2015;10:715–32.
Chu C, Qu K, Zhong FL, Artandi SE, Chang HY. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol Cell. 2011;44:667–78.
Chu C, Zhang QC, da Rocha ST, Flynn RA, Bharadwaj M, Calabrese JM, Magnuson T, Heard E, Chang HY. Systematic discovery of Xist RNA binding proteins. Cell. 2015;161:404–16.
Engreitz JM, Pandya-Jones A, McDonel P, Shishkin A, Sirokman K, Surka C, Kadri S, Xing J, Goren A, Lander ES, Plath K, Guttman M. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science. 2013;341:1237973.
Simon MD. Capture hybridization analysis of RNA targets (CHART). Curr Protoc Mol Biol. 2013;Chapter 21:Unit 21.25.
Quinn JJ, Chang HY. In situ dissection of RNA functional subunits by domain-specific chromatin isolation by RNA purification (dChIRP). Methods Mol Biol. 2015;1262:199–213.
Zhang H, Zeitz MJ, Wang H, Niu B, Ge S, Li W, Cui J, Wang G, Qian G, Higgins MJ, Fan X, Hoffman AR, Hu J-F. Long noncoding RNA-mediated intrachromosomal interactions promote imprinting at the Kcnq1 locus. J Cell Biol. 2014;204:61–75.
Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, Thomas K, Presser A, Bernstein BE, van Oudenaarden A, Regev A, Lander ES, Rinn JL. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci U S A. 2009;106:11667–72.
Niranjanakumari S, Lasda E, Brazas R, Garcia-Blanco MA. Reversible cross-linking combined with immunoprecipitation to study RNA-protein interactions in vivo. Methods. 2002;26:182–90.
Ule J, Jensen K, Mele A, Darnell RB. CLIP: a method for identifying protein-RNA interaction sites in living cells. Methods. 2005;37:376–86.
Licatalosi DD, Mele A, Fak JJ, Ule J, Kayikci M, Chi SW, Clark TA, Schweitzer AC, Blume JE, Wang X, Darnell JC, Darnell RB. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature. 2008;456:464–9.
Konig J, Zarnack K, Rot G, Curk T, Kayikci M, Zupan B, Turner DJ, Luscombe NM, Ule J. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat Struct Mol Biol. 2010;17:909–15.
Martin L, Meier M, Lyons SM, Sit RV, Marzluff WF, Quake SR, Chang HY. Systematic reconstruction of RNA functional motifs with high-throughput microfluidics. Nat Meth. 2012;9:1192–4.
Buenrostro JD, Araya CL, Chircus LM, Layton CJ, Chang HY, Snyder MP, Greenleaf WJ. Quantitative analysis of RNA-protein interactions on a massively parallel array reveals biophysical and evolutionary landscapes. Nat Biotechnol. 2014;32:562–8.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Shechner DM, Hacisuleyman E, Younger ST, Rinn JL. Multiplexable, locus-specific targeting of long RNAs with CRISPR-Display. Nat Meth. 2015;12:664–70.
Acknowledgements
We would like to thank Daniel Thomson for his input during the making and proofreading of the manuscript.
Authors’ contributions
NB and JLVM designed the outline of the paper. JLVM and NB created all the figures. NB, JLVM and MED wrote the manuscript. All authors have read and approved the final version of this manuscript.
Competing interests
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional file
Additional file 1: Table S1
lncRNAs and references for Fig. 2. (PDF 106 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Bartonicek, N., Maag, J.L.V. & Dinger, M.E. Long noncoding RNAs in cancer: mechanisms of action and technological advancements. Mol Cancer 15, 43 (2016). https://doi.org/10.1186/s12943-016-0530-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12943-016-0530-6